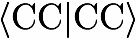
|
Main /
Molecular Geometry InputThe Z-Matrix input is the default choice; use of Cartesian coordinates necessitates an additional keyword (COORD=CARTESIAN) and is explained at use of cartesian coordinates. The Z-matrix assigns a line to each atom in which its position is specified relative to three previously specified atoms. The specification includes a length, an angle, and a dihedral angle (all characterized by variable names to which in the following
section in the ZMAT file numerical values are assigned). The number associated with each atom is determined by its position in the Z-matrix. The essentials of Z-matrix construction are probably best illustrated by considering a Z-matrix for a system of four atoms ABCD, not necessarily linked in any particular order. A -----B
C ---- D
The Z-matrix for this system may be chosen as: A The first line in the Z-matrix just contains the atomic symbol of one of the atoms, say A. There is nothing more to specify for the first atom and it is placed at the origin of the coordinate system. The second line specifies the position of the second atom, say B, relative to the first atom (A). Suppose that the distance between B and A is RAB. The second line then contains the atomic symbol B, followed by the number 1 (A is atom number 1) and the distance RAB to this atom.
No further specification is necessary for the second atom and it is then placed on the positive z-axis. For the specification of the third atom, say C, a distance and an angle are needed. Using the distance between atoms A and C, RAC, and the angle CAB formed by atoms C A B (in this order, see picture), the third line is 'C 1 RAC 2 CAB'. An alternative choice would be the use of the distance between C and B and of the angle CBA. No dihedral angle is required, as the third atom is placed in the xz plane. Finally for the fourth atom the full set of bond length, bond angle, and
dihedral angle must be given and might in the present case possibly take the form 'D 3 RCD 2 DCB 1 TAU'.
This line simply states that the fourth atom, say D, has a distance RCD to C, an angle DCB to C and B and a dihedral angle TAU to C,B, and A. The dihedral angle is here defined as the angle between the BCD and ABC planes. For systems with more than four atoms, the fifth and subsequent lines follow the same pattern as the fourth line of the example given above, i.e., they contain a length, an angle, and a dihedral angle together with the number of three previously specified centers. It should be emphasized that this simple example for a Z-matrix works for a tetraatomic molecule such as hydrogen peroxide, but does not suffice for all cases.
For linear molecules (such as acetylene) as well as ring compounds, the use of "dummy" atoms, i.e. simply a point in space with no charge and no basis functions, is essential. Definition of dihedral angle: The dihedral angle for the four atoms ABCD as specified in the example
above is defined as follows: It is the angle needed to rotate the projection of the AB vector (in a plane with the BC vector as normal vector) into the projection of the CD vector into the same plane. Anti-clockwise rotation with respect to BC as rotation axis is taken as positive; the values are restricted to -180 to 180 degrees. The concept of a dihedral angle is best illustrated by the following picture 1
/
/
2 dihedral angle 4 3 2 1
3
4
It is also useful to remember that a 'cis' orientation corresponds to a dihedral angle of 0 degrees, an 'orthogonal' orientation to either -90 or 90 degrees, and a 'trans' orientation to a dihedral angle of 180 degrees. Dummy atoms and linear molecules As bond angles have to be between 0 and 180 degrees thereby excluding the two limiting cases, the specification of a linear arrangement of three atoms is somewhat tricky in the Z-mat notation. However, it is possible by using dummy atoms (specified by X in the Z-mat) which just represent arbitrary points in space. For the HCN molecule, the Z-Mat can be easily specified using a dummy atoms as follows C which is explained via the corresponding picture. picture of HCN with dummy atoms The dummy atom is places with an angle of 90 degrees with respect to the CN bond. In this way is it possible to place the hydrogen atom on the z-axis by specifying an angle of 90 degrees with respect to the dummy atom together with a dihedral angle for HCXN of 180 degrees. Note that dummy atoms (X) just represent arbitrary points in space which are used to simplify the construction of the Z-matrix. They do not carry a charge nor will be a basis set attached to them. Ghost atoms Ghost atoms, which are specified by the symbol GH have zero nuclear charge. However, unlike dummy atoms ghost atoms serve as a center for basis functions. This feature is particularly useful for calculations performed to determine the basis set superposition error (BSSE) and has several other potential applications such as describing lone pair electrons of a molecule by additional functions not centered at any of the molecular nuclei. Note that symmetry can be used in calculations involving ghost atoms. However, the used symmetry is in these cases restricted to the symmetry of the supermolecule made up of the real AND ghost atoms. Depending on the chosen ghost atoms, the corresponding point group might be a subgroup of the actual point group of the real molecule. Note that in calculations involving ghost atoms, the basis set has to be supplied in all cases explicitly via the BASIS=SPECIAL feature explained in section X.XX. Furthermore, it should be noted that currently only single point calculations are possible. example for bsse calculation .... TDA, pairs of dihedral angle ... symmetry usage example H2O ZMAT for water (H2O) NH3 ZMAT for ammonia (NH3) N 1 with D120 as 120 degrees CH4 ZMAT for methane (CH4) with TDA as the tetrahedral angle (109.4712206... degrees) H2O2 ZMAT for H2O2 HCN ZMAT for hydrogen cynide (HCN) with the angles set A90 equal to 90 degrees and D180 to 180 degrees C2H2 ZMAT for acetylene (C2H2) with angles A90 equal to 90 and D180 equal to 180 degrees C6H6 ZMAT for benzene (C6H6) with angles A60 equal 60 and D180 equal to 180 degress. RCC denotes here the CC distance and RXH the sum of CC and CH distance. Variable specification The second part of the Z-matrix contains the definition for the used variable names in the sense that values for the used internal coordinates are specified. This part just contains of a list with one line for each variable, e.g. PNM=value where PNM is one of the coordinates used in the Z-matrix and value is the value assigned to this coordinates. The parameters may be given in any order. Angles must be enters in degrees, distances either in Angström (default) or atomic units (together with the keyword UNITS=BOHR). Bond angles of 0 and 180 degrees are not allowed, since these lead to a singularity in the transformation between Cartesian and internal coordinates (for the treatment of linear molecules, see above).
An example is the Z-Matrix of methane in which also the tetrahedral angle TDA is used Z-matrix for methane The end of the first part of the Z-matrix is indicated by a blank line. in the given example, the internal coordinate R (actually the CH distance) a value of 1.085 A is assigned, while for the angle TDA (actually the HCH angle) a value of 109.471 degrees is specified and internally replaced by the correct value. For the dihedral angle D120 a value of 120 degrees is given. Specification of coordinates to be varied in geometry optimizations In the case of geometry optimization, an asterisk * immediately after the variable name in the Z-matrix indicates that the particular internal coordinate is to be optimized. For methane, for example, the Z-matrix then should be as Z-matrix for methane as only the CH distance R need to be optimized. The tetrahedral angle as well as the dihedral angle D120 are assumed to keep their initial values during the geometry optimization. It should be noted that via the file JMOL.plot or MOLDEN (created by the module xjoda) a visualization of the geometry of the molecule is possible by using
JMOL (geometries) or MOLDEN (geometries, molecular orbitals, electron densities). |